Here we will discuss about all parameters used in Qsar study. Here parameters which are used in 2D and 3D both are discussed here. The selection of compound having following criteria mean it have to follow Lipinski’s Rule of 5.The Lipinski’s Rule of 5 as follows13:
Have 5 or fewer H-bond donors (expressed as the sum of OHs and NHs)
Have a MW <500
Log P less than 5
Have 10 or less H-bond acceptors (expressed as the sum of Ns and Os)
Molecular descriptors are numerical values that characterize properties of molecules. Molecular descriptors encoded structural features of molecules as numerical descriptors. Vary in complexity of encoded information and in compute time Examples: 1) Physicochemical properties (empirical)2)Values from algorithms, such as 2D fingerprints15.
2D descriptors only use the atoms and connection information of the molecule. Internal 3D descriptors use 3D coordinate information about each molecule; however, they are invariant to rotations and translations of the conformation. External 3D descriptors also use 3D coordinate information but also require an absolute frame of reference (e.g., molecules docked into the same14.
Here I try my best that student can easily understand various parameters used in Qsar. In QSAR analysis, it is imperative that the biological data be both accurate and precise to develop a meaningful model15. It must be realized that any resulting QSAR model that is developed is only as valid statistically as the data that led to its development16.
The equilibrium constants and rate constants that are used extensively in physical organic chemistry and medicinal chemistry are related to free energy values DG18. Thus for use in QSAR, standard biological equilibrium constants such as Ki or Km should be used in QSAR studies. Likewise only standard rate constants should be deemed appropriate for a QSAR analysis17.
There are many physical, structural, and chemical properties which have been studied by the QSAR approach, but the most commonly studied are hydrophobic, electronic, and steric33. This is because it is possible to quantify these effects relatively easily19. In particular, hydrophobic properties can be easily quantified for complete molecules or for individual substituents. On the other hand, electronic and steric properties are more difficult to quantify, and quantification is only really feasible for individual substituents. The three most studied physicochemical properties will now be considered in some detail20.
Hydrophobicity:-
The hydrophobic character of a drug is crucial to how easily it crosses cell membranes and may also be important in receptor interactions21. Changing substituents on a drug may well have significant effects on its hydrophobic character and hence its biological activity. Therefore, it is important to have a means of predicting this quantitatively22.
The partition coefficient (P)
The hydrophobic character of a drug can be measured experimentally by testing the drug's relative distribution in an octanol/water mixture22. Hydrophobic molecules will prefer to dissolve in the octanol layer of this two-phase system34, whereas hydrophilic molecules will prefer the aqueous layer. The relative distribution is known as the partition coefficient (P) and is obtained from the following equation:
Concentration of drug in octanol
Concentration of drug in aqueous solution
Hydrophobic compounds will have a high P value, whereas hydrophilic compounds will have a low P value.
Biological activity normally expressed as 1/C, where C = [drug] required to achieve a defined level of biological activity. The more active drugs require lower concentration35.
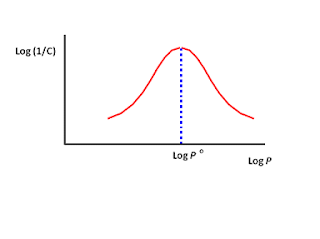
The graph is drawn by plotting log (1/C) versus log P. The scale of numbers involved in measuring C and P usually covers several factors of ten and so the use of logarithms allows the use of more manageable numbers22.
(Figure 1.5.1 parabolic curve)
Here, the biological activity increases as log P increases until maximum value is obtained. The value of log P at the maximum (logP0) represents the optimum partition coefficient for biological activity. Beyond that point, an increase in LogP results in a decrease in biological activity36.
In situations where the partition coefficient is the only factor influencing biological activity, the parabolic curve can be expressed by the mathematical equation:
2-k2 logp+k3
The (LogP) 2 terms has a negative sign in front of it. When P is small, the (LogP) 2 terms is very small and the equation is dominated by the LogP term. This represents the first part of the graph where activity increases with increasing P.
If graph is extended to very high log P values, then get a parabolic curve. Reasons:1) poorly soluble in aqueous phase 2)trapped in fat depots
It is more susceptible to metabolism.
The substituent Hydrophobicity constant (π)22:-
In order to get P we have to measure it experimentally and that means that we have to synthesize the compounds. It would be much better if we could calculate P theoretically and decide in advance whether the compound is worth synthesizing. QSAR would then allow us to target the most promising looking structures.
For example, if we were planning to synthesize a range of barbiturate structures, we could calculate log P values for them all and concentrate on the structures which had LogP values closest to the optimum logP0 value for barbiturates. Fortunately, partition coefficients can be calculated by knowing the contribution that various substituents make to Hydrophobicity37. This contribution is known as the substituent hydrophobicity constant.
The substituent hydrophobicity constant is a measure of how hydrophobic a substituent is, relative to hydrogen. The value can be obtained as follows. Partition coefficients are measured experimentally for a standard compound with and without a substituent (X). The hydrophobicity constant (TTx) for the substituent (X) is then obtained using the following equation:
X = logPx - LogPH
Where PH is the partition coefficient for the standard compound, and Px is the partition coefficient for the standard compound with the substituent. A positive value of indicates that the substituent is more hydrophobic than hydrogen. A negative value indicates that the substituent is less hydrophobic38.
Example:
Here are some value are given. These are considering in calculating parameter. For example methyl has value 0.52 .these are approx.value.in different book value are somewhat changed on point value.
It should be noted that values for aromatic substituents are different from those used for aliphatic substituents.
1) Benzene: - Log P = 2.13
2) Chlorobenzene: - Log P = 2.84
Steric factors22:-
Quantifying steric properties is more difficult than quantifying hydrophobic or electronic properties. Several methods have been tried and three are described here. It is highly unlikely that a drug's biological activity will be affected by steric factors39.
1) Taft's steric factor
Attempts have been made to quantify the steric features of substituents by using Taft's steric factor (Es).The number of substituents which can be studied by this method is restricted.
Es=log (K/Ko) A
2) Molar refractivity (MR)
Molar refractivity is a measure of the volume occupied by an atom or group of atoms. It is given by following equation.
Where n is the index of refraction, M W is the molecular weight, and d is the density. The term MW/d defines a volume, while the (n2 — l)/ (w2 + 2) term provides a correction factor by defining how easily the substituent can be polarized; this is particularly significant if the substituent has pi electrons or lone pairs of electrons.
3) Verloop steric parameter
Another approach to measuring the steric factor involves a computer programme called STERIMOL which calculates steric substituent values (Verloop steric parameters) from standard bond angles, van der Waals radii, bond lengths, and possible conformations for the substituent.
Here various parameter used in 3D Qsar given. They are like electronic, thermodynamic and steric parameter. These are some of parameter. In 3D thousand parameters are calculated.
1) 3D Electronic Descriptor23:
Polarizability: The Polarizability property provides information about the distribution of electrons based on presence of an applied electric field. Polarizability data is often used in other equations for evaluation of optical properties of molecules.
Dipole movement: The dipole moment is the first derivative of the energy with respect to an applied electric field. It measures the asymmetry in the molecular charge distribution and is reported as a vector in three dimensions.
HOMO Energy: Energy of the highest occupied molecular orbital.
LUMO Energy: Energy in of the lowest unoccupied molecular orbital.
2) 3D Thermodynamic descriptors23:
Boiling point: The boiling point for the structure at 1 atmospheric pressure.
Charge-Charge Energy: The sum of the electrostatic energy representing the pair wise interaction of charged atoms.
Charge-Dipole Energy: The sum of the electrostatic energy terms resulting from interaction of a dipole and charged species.
Critical Temperature: The temperature (Tc) above which the gas forms of the structure cannot be liquefied, no matter the applied pressure.
Critical Volume: The volume occupied (Vc) at the compounds critical temperature and pressure.
LogP: The logarithm of the partition coefficient for n-octanol/water.
Melting Point: The melting point for the structure at 1 atm.
Standard Gibbs free energy: The Gibbs free energy (DG) for the structure at 298.15 K and 1 atm.
Vapor Pressure: The vapor pressure for the structure at 25° C.
Dipole-Dipole Energy: The sum of the electrostatic energy terms resulting from interaction of two dipoles
Stretch-Bend Energy: The sum of the stretch-bend coupling terms of the force-field equation.
Total Energy: The sum of all terms the force-field equation
Van der Waals Energy: The sum of pair wise van der Waals interaction energy terms for atoms
3) 3D Steric Descriptors23:-
Exact Mass: The exact molecular mass of the molecule, where atomic masses of each atom are based on the most common isotope for the element.
Molecular Formula: The molecular formula showing the exact number of atoms of each element in the molecule.
Molecular Weight: The average molecular mass of the structure, where atomic masses are based on the weighted average of all isotope masses for the element.
So, these are various parameters of 2D and 3D .All are given in detail. The value of various parameters is given in various books like burger, history of Qsar by C.D.SELLASSIE and in many books values are given.



0 comments:
Post a Comment